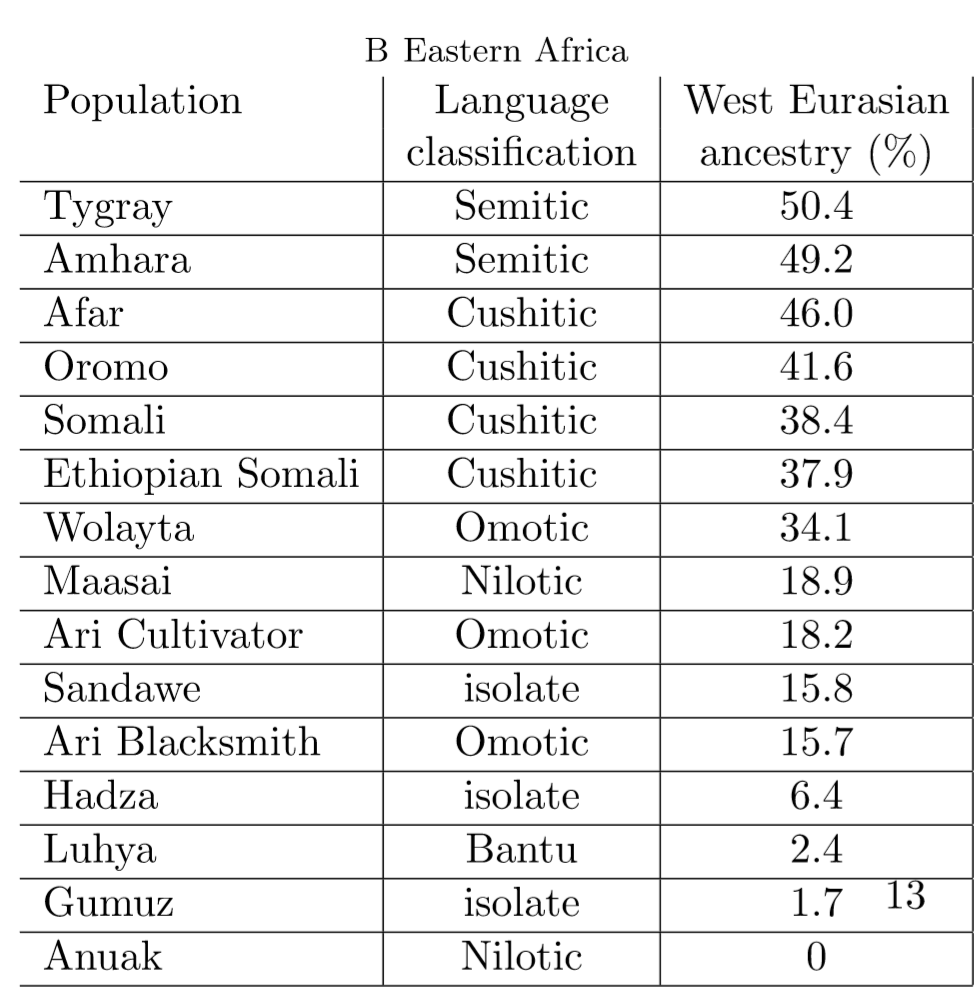
I recently encountered a member on a Forum I frequent and he made mention of Hodgson et al.'s following statements in its abstract which is called back to on wikipedia when discussing Somali autosomal DNA:
"The non-African ancestry in the HOA,
which is primarily attributed to a novel Ethio-Somali inferred ancestry
component, is significantly differentiated from all neighboring
non-African ancestries in North Africa, the Levant, and Arabia. The
Ethio-Somali ancestry is found in all admixed HOA ethnic groups, shows
little inter-individual variance within these ethnic groups, is
estimated to have diverged from all other non-African ancestries by at
least 23 ka, and does not carry the unique Arabian lactase persistence
allele that arose about 4 ka."
Along with its following claim that this supposed component is related to the "Maghrebi" component:
"A single prehistoric migration of both the Maghrebi and the Ethio-Somali
back into Africa is the most parsimonious hypothesis. That is, a common
ancestral population migrated into northeast Africa through the Sinai
and then split into two, with one branch continuing west across North
Africa and the other heading south into the HOA. For the Ethio-Somali,
the lowest FST value from the ADMIXTURE estimated ancestral allele frequencies is with the Maghrebi (Text S1), which is consistent with a common origin hypothesis."
I often wonder though how some of these studies make it onto journals or how they were made by professionals. Ultimately, "Ethio-Somali" is not a real ancestral component, nor is it truly related to another "not real" component they dub "Maghrebi" (it is but not in the way they perhaps posit). The study does good work in some respects such as showing us the flaws in dating systems for admixture such as ALDER & ROLLOFF. Horners used to and sometimes still do have their admixture dated to about 2 to 3 thousand years ago using dating methods and of course those dating points make no sense and are generally not possible. They also in my own opinion do a good job in dating some of the admixture in the Horn as being newer than some of the rest; ultimately identifying the newer Middle Eastern input into Agaws ("Afar" samples are in fact Xamtanga speaking Agaws) & Ḥabeshas (i.e those who have been tested so far such as Tigrinyas & Amharas) over Somalis but otherwise...
It's results are honestly based on finding some mixed component at K=12 via the ADMIXTURE software and by some insane leap; it assumes that the component is not mixed/ "Near Eastern/ Middle Eastern + East African/ "Nilo-Saharan" " as it really is if you simply observe their own run:
Like in most ADMIXTURE based runs using African & Eurasian (Out of Africa) populations; the run at K=2 starts out with every population being utilized coming out as either "Eurasian" or "African" (in this case these two macro-components are represented by "European" & "Niger-Congo") and observe the Horn of Africa populations. We all look like a clear mix of the two with Somalis looking almost half-half but visibly more "African" shifted than "Eurasian" shifted yet Ethio-Somali makes up a whooping 57%~ of the ancestry in them and it is supposedly 100% Non-African?
Want further proof? I believe some old posts I made on that Forum I frequent will be adequate:
" Ethio-Somali is similar to the Maghrebi component in that it is slightly "mixed". It has an ancient East African
influence however that influence is minor, I suppose and it is
predominantly a West Eurasian component that is most probably a very
ancient variant of Southwest Asian/ "Arabian". You can observe it's clear "Arabian" like affinities at the lower Ks in the Hodgson study.
Observe K=6 in that image where all the ancestry in the Horn is seen
entirely as "Arabian" (dark brown) and "Nilo-Saharan" (light blue).
There is some "Ethiopic"/ Omotic ancestry hidden for all Horners in that
Nilo-Saharan but then I want you to look at K=12 in that image where
all of a sudden Horners seem to be pred "Ethio-Somali" especially in the
case of Somalis yet there's less Ethiopic (purple) + Nilo-Saharan than
there was "Nilo-Saharan" at K=6. I have other ways to show you it's
influences so just ask."
"Maghrebi" which is another "component" the study finds is also not really real... Now, any respectable geneticist who's studied the populations of the Maghreb/ Northwest Africans (Imazighen/ Berbers) is aware that they are "African" admixed at a rate of about 20 to 30%~. In fact the very study Hodgson et al. gets its Horner samples from notes the African admixture in Maghrebis as do various other inquiries into Maghrebi ancestry such as a study from 2012 titled "Genomic Ancestry of North Africans Supports Back-to-Africa Migration" [2] [3].
Even independent ADMIXTURE runs created by various individuals who are not even geneticists with degrees under their belt can obviously notice as much [-] , heck; a meager & intensely flawed profit orientated ancestry pinpointing service like FTDNA managed to notice the African admixture in a random Tunisian.[-] In terms of having African admixture; the same can be said for Peninsula Arabians, Egyptians and the like, the former of which having their admixture confirmed by not only Hodgson et al. but various other runs such as the recent revolutionary study focused on ancient genomes by Losif Lazaridis. [1] [4]
To be fair it does note the African admixture in most Maghrebis outside of the "Maghrebi component" however it finds that "Tunisians" (this is more of a genetically isolated Tunisian population (supposedly practicing endogamy) that may not entirely be representative of all Tunisians [3]) are more or less entirely made up of this supposed ancestral component which they claim diverged from Ethio-Somali (according to them-> a 100% Non-African component) but obviously from looking at these Tunisians at K=2 in their own analysis one can see the African influence in the component also & at the higher Ks before it's introduction-> various influences such as "Arabian", "European" (more representative of the more "Northern" admixture of the Middle East. I.e. a component often dubbed "Mediterranean" in this case).
In a PCA (Principal Component Analysis), these Tunisians don't even cluster different from other Maghrebis with only likely perhaps the most African admixed such as some Southern Moroccans and a few outliers here and there breaking off noticeably:
Yet these Tunisians are supposed to have a supposedly fully "Eurasian" ("Non-African") ancestral component that makes up pretty much all of their ancestry but they cluster relatively no differently from Moroccans, Sahrawis, Algerians, Libyans and Mozabites who have very visible post-formation of this Magrebi component Niger-Congo & "Nilo-Saharan" (what I often call AEA/ Ancestral East African) admixture. These Tunisians have been tested in various ADMIXTURE runs like those independent ones and have been found to be about 10 to 20%~ African admixed. If you don't trust them (I don't know why... Some of them clearly know more than this study does in terms of ADMIXTURE components) then really just know that the same study that identified those Tunisians' isolated-ness on a genetic level, also pretty much finds them to be like other North Africans to be African admixed. [3]
For good measure, here is another study that finds similar admixture in them whilst also using those Tunisians who are supposedly according to Hodgson et al. (a study that lets its own ADMIXTURE runs and what they show at the lower Ks as well as its PCA plots go way over its own head) 100% Non-African/ composed of a component related to a supposed 100% Non-African component ("Ethio-Somali"):
"Consequently, genome-wide diversity show similar
patterns with admixture tests suggesting North Africans are a mixture of
ancestral populations related to current Africans and Eurasians with
more affinity towards the out-of-Africa populations than to sub-Saharan
Africans."
They're basically saying what even complete layman should know about Maghrebis/ Berbers/ Imazighen and those Tunisians-> they're a predominantly ancient/ pre-historic Near Eastern population with non-negligible African admixture.
The suggestion that this Maghrebi component is a fully Eurasian/ Out-of-Africa (OoA) component component is utterly laughable and it hence is shocking that this study headed by actual geneticists with degrees claims it is and is pretty much related to an even more African admixed component (30%~ or so based on more accurate estimates of the admixture in Somalis that I will share later in this post from other studies including a more recent one) like Ethio-Somali.What's even more puzzling is that they're aware of and acknowledge what other papers say about "Maghrebi":
"Like the Ethio-Somali, the Maghrebi IAC in North African populations derives from a early back-to-Africa migration.
Studies of North African populations reveal a complex layered history
of admixture in North Africa, with an inferred pre-Last Glacial Maximum
settlement of North Africa by a non-African population followed by gene
flow from European, Middle Eastern, and sub-Saharan African populations
dating from the end of the LGM to the recent past."
[1]
However they still claim it diverged from a component they believe to be Non-African entirely...
The admixture in the Horn ultimately though does show a strong affinity for the admixture in Peninsula Arabians (Saudis, Yemenis, Qatari Arabs, Omanis et al.) which is demonstrable enough at the lower Ks where the component looks more like "Arabian" in Hodgson et al.'s ADMIXTURE run that anything else in the MENA region (Middle East & North Africa):
This fact has been noticeable to other sources like those independent types I shared who also identify the exact same "Arabian" component that Hodgson et al. does however they dub it "Southwest Asian" as I often do, they find far less of the "Mediterranean" (the majority of the ME admixture in Maghrebis) & "Caucasus"("Eurasian" in Hodgson et al. analysis) component in Horners at large. However, interestingly; in terms of uniparental markers (Y DNA + mtDNA markers), Horners do not entirely fit well with Peninsula Arabians however modern Peninsula Arabians are by no means to be mistaken for the pre-historic peoples who mixed with AEA rich East Africans episodically to create us. I say episodically because both in terms of uniparentals and autosomal DNA data like that found through ADMIXTURE-> the admixture interactions that created Horners seem to have been episodic in nature/ we didn't become the way we are in one fell swoop.
To its credit; Hodgson et al. notes the episodic nature of our admixture and of course manages to notice the newer ADMIXTURE in Agaws & Ḥabeshas and many Oromos which I applaud it for doing as this is somewhat useful. Otherwise it's a crap study in that it makes too much of several mixed components it finds at the higher Ks/ it's ADMIXTURE software spits out for it. There have been "Cushitic components" (it's a pseudo-Cushitic component) in the past and even after Hodgson et al. One in Tishkoff et al. in 2009 and one just recently during this month by a study with some really ridiculous and incorrect notions that are heavily disproved by ancient DNA/ genomes like those used in Lazaridis et al. , Fu et al. , Gamba et al. & Raghavan et al. which are really making amazing strides in re-defining Modern Human population genetics. [6] [7]
Just to entertain you, I'll go into that more recent study's findings on Horners... They essentially claim that a component dominating the ancestry in Somalis is a component called "Lowland East Cushitic" and that it is of a "Macro-related-group" with components it dubs "Indian", "European", "Levantine-Caucasian" & "Arabian" and it claims (this is the punch line now) that this component diverged from them 61,000 years ago.
Don't take my word for it:
"Lowland East Cushitic ancestry diverged from the
Caucasian/European/Middle Eastern/south Asian cluster 2,041 generations
or ~61,000 years ago"
[8]
They ultimately find something very much like "Ethio-Somali which they also say makes up not all but a great part of the ancestry in ethnic Somalis"- :
-then they make the humorous claim that it's essentially 100% Non-African by claiming it diverged from "Indian" for example but there is not at all one Indian component, in fact one of the people involved in Lazaridis et al. found 5 years ago what many already know now; that South Asians are predominantly a mixture of a sort of "North Indian" component that is seemingly at least predominantly West Eurasian alongside another component often called "South Indian" which is a component that is clearly differentiated from West Eurasian ones and also seemingly East Eurasian ones as well. [9]
European as well have for a long time been divided into "North" & "Southern" European (autosomal DNA wise) but of course anyone who has been keeping up with European genetics knows how mixed Europeans are based on the findings of Lazaridis et al. , Gamba et al. and hell even the sequencing of Ust-Ishim's genome. These studies are 100% more reliable than this farce and essentially use ancient remains from as far back as 45,000 years ago in Fu et al.'s case for Ust-Ishim and as far back as 24,000 years in the case of MA-1 & Lazaridis' use of his genome for example. They are essentially what is largely accepted in genetic circles now and are truly redefining population genetics.
This study essentially like this dendrogram for the divergence of components demonstrates claims that "Lowland East Cushitic" diverged from those components I mentioned (Oh, look, there's a Berber/ Magrebi component too and look where they claim it sits):
.
Apparently it makes absolutely nothing of its own dendrogram for fst distance between the ancestral components it identifies:
Funny how in terms of fst distance (i.e. distance between these components in a sense): "Lowland East Cushitic" is closer to Nilo-Saharan, Niger-Congo and Omotic than it is to those components it supposedly diverged from 61,000 years ago (a completely laughable and fictutious time frame btw).
.
Oh, Hodgson et al. showed similar fst distance results too btw but chocked it up to the component's supposed 23,000 year divergence from other Non-African components like MENA ones such as Maghrebi being obviously a very long time ago:
Once again it's a supposed Non-African component that's visibly much closer to the African components than any other Non-African component is...
I'm a bigger fan (sarcasm here) of this newer study's ("Shriner et al.") explanation for its pull toward the African components and the well-known pull West Eurasians who lack any known African admixture such as Europeans display toward the African groups in comparison to how East Eurasians (Han Chinese, Onge etc.) pull toward them which is visibly always much farther away.
It's according to Shriner et al. that there were two Out of Africa migrations... The second marking an occasion where the Indian, Kalash, European, Levantine-Caucasian, Berber & Arabian components left Africa after theoretically diverging from Lowland East Cushitic.
This is incredulous fiction and if they'd read a single paper on the latest finds in the genetic field pertaining to what we're learning from ancient genomes finally being sequenced (a thousand times more reliable than studying modern populations to explain pre-historic genetic history)-> they'd know that the reason West Eurasians like Europeans pull towards Africa more is due to Basal Eurasian (whatever the component might ultimately prove to be once we get ancient DNA from the Middle East and hopefully also East Africa sequenced) as Lazaridis et al. and other studies lately have noticed. Fu et al. also proves that the reason modern Europeans are not as close to Ust-Ishim whose genome sits between pre-historic non-Near Eastern admixed (and thus non-Basal Eurasian admixed) West Eurasians like the Western European Hunter-Gatherer dubbed Loschbour & East Eurasians is very likely explainable by something with the affinities Basal Eurasian possesses:
But ultimately, the study (Shriner et al.) is crippled heavily by what the author of Eurogenes describes below:
"The authors are under the illusion that running enough samples with the
ADMIXTURE software can reveal pure ancestral components from thousands
of years ago, and thus uncover the story of the peopling of the world in
all of its glory. That's an incredibly naive view, to put it mildly."
Well said... Incredibly naive, indeed. They have no regard for ancient genomes and just think they can describe Human pre-history via the ADMIXTURE software and whatever it craps out for them. ADMIXTURE software pushes out mixed components like this often, just like it did with Maghrebi for Hodgson et al. and that's really all Ethio-Somali is (beyond perhaps being useful for marking migrations and gene flow from Cushitic or Cushitic-like groups into other groups); a mixed component the software just shat out for Hodgson et al. and they just ate it up.
More accurate results for the admixture in the Horn of Africa is shared for example by Pickrell et al. :
.
The study went so far as to attempt to model the admixture in the Horn of Africa with reference populations and found the admixture to be good fits for models such as the following:
Anuaks, South Sudanese etc. carry the AEA/ Nilo-Saharan component/ it's the majority of their admixture so they indeed if you discount their Niger-Congo admixture are a good fit for the admixture in the Horn.
These results are honestly corroborated even more by PCA plots (Principal Component Analysis) such as even the one shared by Hodgson et al. :
.
or the one shared by Pagani from which Hodgson et al. got its Horner samples from:
.
In fact, every PCA plot/ cluster with Horners present displays that Ethiopian Semitic speakers and Agaw/ Central-Cushitic speakers have a somewhat noticeably greater pull towards "Eurasians" than Somalis, Wolaytas and certain Oromos do. Which is generally in line with Pickrell's findings where they have somewhat more Middle Eastern input than Somalis or Wolaytas (with Somalis having somewhat more than Wolaytas).
These results are further corroborated by a very recent & very thorough study (Gudrasani et al.) on African genetics where the authors use three different methods to identify Eurasian admixture in the African continent and they in Methods 1 & 2 find pretty much the same results Pickrell did with only somewhat more heightened Eurasian admixture among the Horner populations via Method 3:
The Horn populations are not nearly as pre-historic Middle Eastern to Non-African admixed as Hodgson et al. or Shriner et al. posit and the proof is not only in their own data (i.e. Lower Ks of admixture runs, fst distance etc.) but also various other studies such as with Shriner et al. which has proved extremely unreliable if one has even remotely become familiar with studies on ancient genomes which completely disprove its assertions.
Horners generally cluster the way populations with their levels of admixture do. Amharas & Tigrinyas being practically 50%~ Middle Eastern influenced tend to cluster as perhaps near pefect intermediates between the populations of West Asia & populations such as the South Sudanese and Niger-Congo speakers (perhaps with just a slightly greater pull towards the South Sudanese/ Nilotic populations) and Somalis show just an ever so slightly greater pull toward the South Sudanese/ Nilotic populations as they are just practically 40%~ Middle Eastern influenced in contrast and about 60-62%~ East African/ "Nilo-Saharan". Hodgson et al. apparently thinks we should plot at the very fringe of the Maghrebi populations or almost with them given the practically 60%~ Non-African score it gives us based on its mixed component, also given that Maghrebis are essentially 20-30%~ African (they do indeed admit the admixture levels in Moroccans & the like, but merely do not notice the admixture in those isolated Tunisians).
On another note; other studies including Pagani et al. & Gudrasani et al. often do come up with the sort of mixed components of the type that Ethio-Somali & Maghrebi are. I.e. Pagani et al. at the higher Ks in its run could be said to have found a mixed component in the Hadza:
Observe the Hadza in the visual image of everyone's admixture; notice tht they're almost entirely made up of an orange component. The Hadza in truth are mixed and even Gurdasani et al. finds similar results in them but still notes their mixed nature:
They look like a population made up by only one component (or predominantly by one component) however in Gudrasani et al.'s three runs for Eurasian ancestry they come out much like they do in Pickrell et al. with relatively 5 to 6%~ Eurasian admixture (particular pre-historic Middle Eastern admixture) and of course the rest of their components are ones like AEA (Ancestral East African/ "Nilo-Saharan"), as well as certain varieties of Khoisan and "Pygmy".
Luckily unlike Hodgosn et al. or Shriner et al.; studies like Pickrell et al. & Gudrasani et al. had the good sense to note the mixed nature of the populations who came up with such dominant components at the higher Ks via further analysis. In fact the components these "not real" components are made up of are not entirely "pure" in any sense. Southwest Asian or Mediterranean are ultimately (as we've learned via ancient genomic tests on the Neolithic Near Eastern farmers of Europe) mixed components as well in the sense that they seem to be some sort of interaction between West Eurasians somewhat similar to the WHGs/ EFs (Western European Hunter-Gatherers/ European Foragers) of Europe and what Lazaridis et al. and Fu et al. believe maybe a "Basal Eurasian" population; a population that did not participate in the forming of an "autosomal clade" between other Eurasians. I.e. between the ancestors of unmixed West Eurasians like Loschbour and the ancestors of East Eurasians (Han Chinese, Onge etc.)-> basically "Out of Africa isolates" who broke from other OoAs early on; at least that's the running theory for now.
Even Nilo-Saharan/ Ancestral East African as I call it; demonstrates a certain amount of mixture with other divergent African populations via the mtDNA of the populations it peaks in who demonstrate L0 markers for example whilst actually at the lower Ks demonstrating a small amount of Khoisan/ Khoesan admixture:
Simply observe populations with a substantial amount of the "Nilo-Saharan" component at everything from K=3 to K=5 (before the the "Nilo-Saharan"/ AEA component is introduced) and the small amount of Khoisan admixture they show... As brought to my attention by a friend; one of the people who worked on Lazaridis et al. said they might eventually be publishing a paper that confirms the idea that Niger-Congo speakers as their uniparental markers (Y DNA + mtDNA markers) [13] [14] [15] imply; are made up of some divergent & perhaps likely pre-historic African populations [-]. Most of the components that make up "Ethio-Somali" themselves will very likely be thoroughly redefined once we have ancient DNA from the Middle East & East Africa that fits with the sequenced.
Ancestral Components like "Ethio-Somali" take this up a notch though by being a mixture of these much more ancient mixtures (this is ultimately what makes it "not real"/ "less real"/ it is certainly not some "pure" component, especially not a Non-African one). The populations that form components like it often do because they've been isolated for some time (i.e. those Tunisians) and then create in a sense; their own distinct cluster. At least that is the case at times...
It is not a "real component" and the populations of the Horn are by no means 60%~ or more Non-African/ Near Eastern but it is correct that there is non-negligible admixture in the horn with an ultimate and likely very pre-historic origin in the Middle East-> the study doesalso have some ups that I took the time to note.
Reference list:
Notes:
1. It's humorous to note that mere layman sources like the Harappa ancestry project for one actually notice the same admixture levels posed by Gudrasani et al. & Pickrell et al. whilst being awfully close to the admixture levels posited by Pagani et al. (the very study Hodgson et al. gets its samples from). Harappa, Dodecad, MDLP etc. all note about 35 to 40%~ Middle Eastern admixture in Somalis for example and about 48 to 51%~ Middle Eastern admixture in Tigrinyas and Amharas (not entirely sure in the case of MDLP). Funny... Even supposed layman know more than this study does...
2. Make no mistake; Modern Nilotes (Dinka, Anuak et al.) & Peninsula Arabians are by no means an accurate representation of the pre-historic peoples who mixed seemingly episodically to create Horners-> both groups predominantly for one have Niger-Congo admixture (non-negligibly; especially in the case of the Nilotic groups). And of course there's the case of Modern Peninsula Arabians not seeming to be a good fit for the Uniparental markers in the Horn. It's all a little complex but they're just essentially; the closest living populations to those pre-historic peoples.
3. However as a friend brought to my attention, the L0 in the Horn or AEA carrying populations is likely not "Khoisan" admixture despite what shows up at the lower Ks. I'll merely quote him for good measure:
"Also, I wouldn't say AEA has "Khoisan" admixture, since the L0
haplogroup is over 100,000 years old and East Africans carry different
L0 lineages compared to the southern African Khoisan. It's a very
distant relation, and you'll find more recent mtDNA relations shared
with West Africans. Certain L2, L3 sublineages have common ancestry with
West African lineages, and almost all of the L0 in the Horn belongs to
L0a, which is also found in West Africa. Non-L0a L0 lineages are mainly
found south of the Horn."